Advanced human liver models for the assessment of drug-induced liver injury
Article information
Abstract
Drug safety issues continue to occur even with drugs that are approved after the completion of clinical studies. Drug-induced liver injury (DILI) is a major obstacle to drug development, because the liver is the primary site of drug metabolism, and injuries caused during this process are severe. Conventional in vitro human liver models, such as 2-dimensional hepatic cell lines, lack in vivo physiological relevance, and animal studies have limitations in the form of species differences and regulatory restrictions. To resolve this issue, an increasing number of 3-dimensional human liver systems, including organoids, are being developed. In this review, we provide an overview of recent assessments of DILI prediction, approaches for in vitro hepatotoxicity evaluation, and a variety of advanced human liver models. We discuss the advantages, limitations, and future perspectives of current human liver models for accurate drug safety evaluations.
Introduction
Drug development requires complex steps, including drug discovery, in vitro and in vivo nonclinical trials, clinical trials, and the United States Food and Drug Administration (FDA) approval. The entire process typically takes more than 10 years and costs $3 billion [1]. Additionally, the success rate of clinical trials is lower than 10%, and drugs are withdrawn from the market owing to safety issues even after their approval. Drug-induced liver injury (DILI), one of the leading causes of drug development failure, accounts for 18% of all drug withdrawals from the market between 1953 and 2013 [2]. Statistically, only approximately 50% of compounds exhibiting liver toxicity in humans have been identified through animal studies, which have recently been restricted due to ethical concerns [3]. Therefore, more accurate model systems are urgently required to reduce these discrepancies and concerns, as well as to accurately predict human responses.
In this regard, primary human hepatocytes (PHHs) obtained from human liver tissue are considered the gold standard for the evaluation of hepatotoxicity. However, the function of PHHs in vitro rapidly decreases in conventional 2-dimensional (2D) culture formats; therefore, 3-dimensional (3D) culture systems have been extensively studied to overcome this limitation. PHHs in 3D culture maintain their function in vitro for over 2 weeks and exhibit accurate toxicity prediction results compared to those in 2D culture [4,5]. Furthermore, 3D organoids have recently emerged as a novel alternative source for human liver models with advanced native tissue architecture and function [6–8]. As a mechanistic understanding of DILI-related adverse outcomes is lacking compared with other organ-specific toxicities, advanced liver models, including organoids, could be a valuable platform that reflects human responses, and patient-derived liver organoids can also serve as personalized drug testing platforms [9,10]. This review focuses on recent studies on DILI risk prediction and discusses the current status and future perspectives of advanced human liver models.
Ethics statement: This study was a literature review of previously published studies and was therefore exempt from institutional review board approval.
DILI classification
DILI is a rare but potentially fatal cause of liver failure associated with adverse drug reactions [11]. These unexpected outcomes are mainly related to the central roles of the liver in the uptake, accumulation, and metabolism (biotransformation) of xenobiotics (Fig. 1A). There are 2 types of DILI: intrinsic DILI occurs in a predictable and usually dose-dependent manner (e.g., acetaminophen [APAP]), and idiosyncratic DILI occurs infrequently and inconsistently at toxic doses of drugs with varied phenotypes. DILI prediction remains challenging owing to the complexity and uncertainty of the associated reactions. To address these issues, the National Center for Toxicology Research at the FDA established a dataset of FDA-approved drugs that induce DILI, called the Liver Toxicity Knowledge Base. This database contains integrated diverse information related to mechanisms, drug metabolism, histopathology, therapeutic uses, targets, and adverse effects useful for DILI evaluation and prediction [12]. DILI severity is categorized by an 8-level system into 3 groups: severe (levels 6, 7, and 8), moderate (levels 4 and 5), and mild (levels 1, 2, and 3) [13] (Fig. 1B). On analyzing the Drug Induced Liver Injury Rank dataset, including 1,036 FDA-approved drugs, liver aminotransferase (alanine aminotransferase [ALT] or aspartate aminotransferase [AST]) increase, a mild concern, accounted for approximately 30%, followed by the most severe concern, fatal hepatotoxicity, such as death or need for liver transplantation, at 15.3% (Fig. 1B). Despite many efforts to model human DILI, there is no approved model for DILI prediction on a nonclinical drug validation platform. As described below, hepatotoxicity evaluation methods in vitro and advanced human liver models reflecting the critical pathologic features of DILI may be invaluable resources for successful drug development.
In vitro hepatotoxicity evaluation
There are no perfect models to emulate the clinical symptoms of DILI; however, several in vitro hepatotoxicity evaluation methods have been developed (Fig. 2). Most DILI-inducing drugs lead to some form of hepatic dysfunction, such as decreased albumin (ALB) and urea secretion, increased serum ALT or AST levels, and cholestasis due to abnormal bile acid flow [14,15]. Some of these pathological phenotypes were successfully reproduced using PHHs treated with 10 known hepatotoxic drugs [16]. ALB secretion by PHHs was decreased after treatment with all 10 compounds by at least 50%, urea secretion and ATP production were decreased using 9 of these drugs, and glutathione (GSH) was decreased with 7 of these compounds, indicating a high degree of concordance between cellular phenotypes and DILI. In addition, the hepatotoxicity of valproic acid, an anti-epileptic drug, was detected in vitro in the form of a significant dose-dependent increase in ALT and AST in the culture medium [17].

In vitro hepatotoxicity evaluation methods. ALB, albumin; ALT, alanine aminotransferase; AST, aspartate aminotransferase; ROS, reactive oxygen species; OXPHOS, oxidative phosphorylation; BSEP, bile salt export pump.
Mitochondrial toxicity is also a critical mechanism of DILI. Troglitazone, an anti-diabetic medication, was withdrawn in 2000 due to rare but severe hepatotoxicity; significant elevations in serum ALT level and necrosis in liver biopsies were detected with the use of this drug [18]. Quinone-based drugs such as troglitazone and APAP are transformed into reactive metabolites by cytochrome P450 (CYP) enzymes [19]. Their intermediate metabolites lead to the accumulation of reducing equivalents that generate reactive oxygen species (ROS), which cause damage to mitochondrial proteins, lipids, and DNA, and finally induce apoptosis by releasing cytochrome C and apoptosis-inducing factor from the mitochondrial intermembrane spaces [20]. Trovafloxacin, an antibiotic, was also withdrawn from the market in 2000 owing to hepatotoxicity, but the mechanisms of trovafloxacin-induced hepatotoxicity were underdetermined. In a liver organoid model, trovafloxacin inhibited mitochondrial respiration, oxidative phosphorylation (OXPHOS) at doses lower than those that produced peak plasma concentrations in humans [8]. In addition, retrorsine, a phytotoxin that induces developmental toxicity, was also reported to possess mitochondrial toxicant functions that reduce mitochondrial respiration and glycolysis capacity [21]. Mitochondrial toxicity and abnormal cellular metabolism can be determined by measuring mitochondria-specific ROS, mitochondrial membrane potential, OXPHOS (by indicators such as the oxygen consumption rate) and glycolysis (by the extracellular acidification rate), and cytochrome C release in vitro [8]. Cell viability measurements via cell number counting, cell cytotoxicity assays, lactate dehydrogenase release, ATP assays, and cell cycle assays are fundamental options.
Cholestasis is characterized by impaired bile secretion and flow due to direct damage to cholangiocytes and small bile ducts or inhibition of bile transporters in hepatocytes, which leads to hepatocellular retention of bile salts and subsequent cytotoxicity, followed by liver injury. Cholestatic DILI is distinct from hepatocellular DILI by an increase in alkaline phosphatase alone and by its tendency to be chronic [11]. Cholestatic and mixed hepatocellular/cholestatic injuries occur in 20% to 40% of all cases of DILI, and the mortality rate of drug-induced cholestasis (DIC) is as high as 10% [22]. One of the main possible mechanisms of DIC is inhibition of the bile salt export pump (BSEP). Bosentan, a drug used for treating pulmonary hypertension with a boxed warning for hepatotoxicity, and cyclosporine A, an immunosuppressant, are known to induce cholestatic DILI through their potent inhibition of BSEP [23]. Troglitazone sulfate, the major metabolite of troglitazone, is also known to function as a competitive BSEP inhibitor, leading to cholestasis and mitochondrial damage [24]. Drugs that inhibit BSEP are more likely to cause idiosyncratic DILI than those that do not [11]. Cholestasis can be measured in vitro by gene expression of bile acid transporters and functional tests with the treatment of fluorescein-labeled bile acids (Fig. 2). As a novel approach, application studies using advanced in vitro liver models for the precise prediction of DILI are discussed below.
In vitro human liver models
Conventional hepatic cell lines, such as HepG2 and Huh7, have been widely used for in vitro testing owing to their practical proliferative capacity. However, the relevance of the data in humans, including those on drug metabolism, is inferior. PHHs are superior to conventional cell lines in terms of hepatic functions but they have limited viability in vitro; therefore, 3D culture systems have been applied to overcome this limitation (Fig. 3). One such application is the spheroid model, which is a simple 3D cell aggregate [25–27] that consists of only hepatocytes or co-cultures of hepatocytes with non-parenchymal cells, including hepatic stellate cells, sinusoid endothelial cells, and Kupffer cells [28]. In toxicity prediction, spheroids exhibit approximately 100% specificity (a non-toxic compound is rated negative) whereas their sensitivity (a toxic compound is rated positive) is lower than 50% [29,30]. The sensitivity value is significantly higher than that of conventional 2D models, but there are limitations, such as insufficient interactions between cells and the extracellular matrix, difficulty in drug diffusion into the inside of spheroids, and difficulty in long-term culture and expansion [10,31–33].
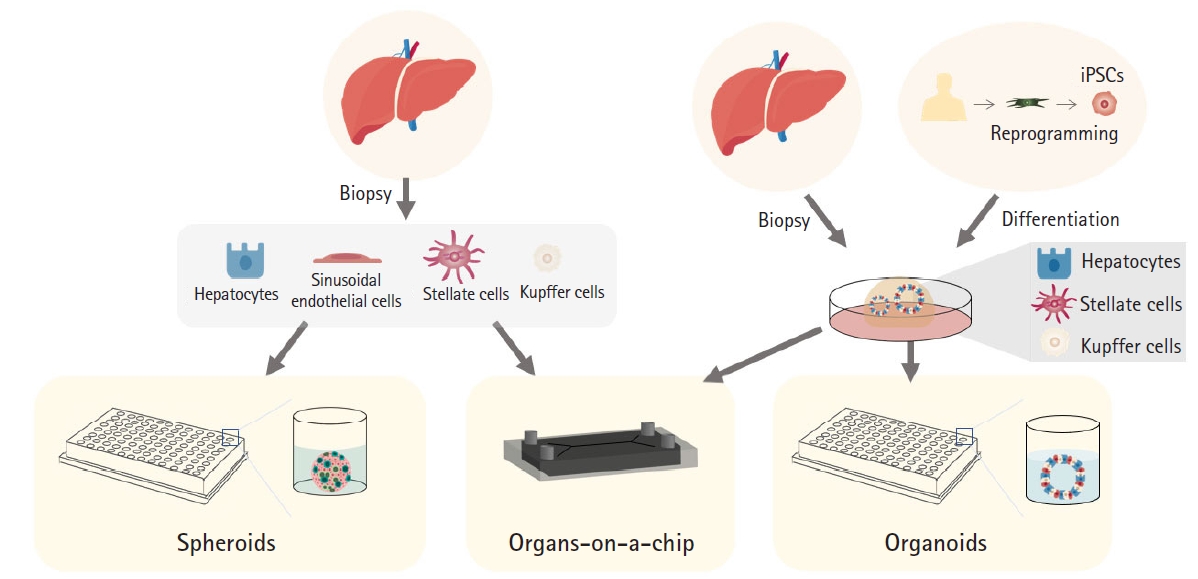
Current in vitro human liver models for drug-induced liver injury prediction. iPSC, induced pluripotent stem cell.
Organs-on-a-chip are also utilized as advanced liver model systems that adapt a multichannel 3D microfluidic platform to emulate the activity, dynamics, and physiological responses of organ systems [32,34]. The oxygen gradient and metabolic zonation can be implemented in a liver chip to predict zonation-specific toxicity [35,36]. Hepatocytes in pericentral zone 3 express high levels of CYP enzymes and execute xenobiotic metabolism. Artificial hepatic zonation via a Wnt gradient represents zone 3-specific APAP toxicity that forms toxic reactive metabolites through CYP2E1, CYP1A2, and CYP3A4 [5,37–40]. Furthermore, inter-organ interactions can be simulated using this bioengineering technology, which mirrors absorption (gut), distribution, metabolism (liver), and excretion (kidney) [34]. However, materials such as polydimethylsiloxane, which are generally used in this model, are vulnerable to non-specific binding, and high-throughput screening is more difficult with this method than with static cultures [29].
Organoids that maintain morphological and functional complexity may be beneficial in DILI risk assessment as they reflect various symptoms of toxicity. Liver organoids can be generated from both human liver tissue biopsies and induced pluripotent stem cells (iPSCs) that provide patient-specific and physiologically organotypic systems that recapitulate organ structure, cell composition, and function [7,41]. Practically, expandable and freezable liver organoids can overcome the cell source limitation problem of PHHs [42]. More importantly, scalability contributes to providing consistent and robust drug evaluation platforms in a reproducible manner. Further, functionally competent liver organoids accurately evaluate toxicity at clinically relevant concentrations of market-withdrawn drugs such as troglitazone and trovafloxacin [8]. DILI phenotypes, including ROS generation, decreased GSH levels, and nuclear structural damage, were distinctly detected by multiplexed high-content screening in liver organoids treated with these drugs. Furthermore, the cytotoxic DILI caused by trovafloxacin and its non-toxic analog, levofloxacin, were clearly distinguished. In addition, the recovery potential of liver organoids was observed in real time after treatment with high-dose APAP and its removal. Therefore, chronic toxicity evaluation is also possible through long-term repeated treatment, which could be another valuable advantage of using organoids in DILI risk assessment [8].
Multi-tissue organoids consisting of parenchymal and non-parenchymal liver cells, including hepatic stellate, sinusoid endothelial, and Kupffer cells, have been generated [6,43]. iPSC-based multi-tissue organoids are composed of all cell types from identical donors; thus, personalized DILI prediction is possible, and better integration of hepatocytes and non-parenchymal cells is expected [43–45]. More importantly, functionally proficient organoids can also reflect complex DILI responses, including inflammation and fibrosis, which mainly occur due to the interactions of hepatocytes with non-parenchymal tissue cells [43]. In addition, cholestasis was predicted with high sensitivity and specificity (88.7% and 88.9%, respectively), based on a hollow-like organoid structure with polarity for bile acid uptake and excretion (Fig. 2). High-throughput screening is also possible using a 384-well plate scale with high fidelity [46]. Finally, applications of “organoids”-on-a-chip are currently under development [47]. Liver organoids have also been adapted to a chip system, enabling enhanced liver function and accurate prediction of DILI for a drug halted at phase 2 clinical trials [48].
Conclusion and future perspectives
DILI remains a major cause of drug safety issues, and hepatotoxicity risk assessment remains difficult due to the potential complications of DILI. Significant progress has been made in the development of physiologically relevant in vitro human liver models and hepatotoxicity evaluation methods [49]. Notably, with advances in stem cell research and 3D tissue engineering, advanced liver models such as organoids have been applied for DILI prediction. Although organoids have been a novel alternative source for human-relevant model systems with respect to their architecture and multiple cell type compositions, their critical levels of baseline performance in drug metabolism and hepatocyte function have not fully reached the levels achieved with PHHs. Technological developments for the enhancement of the mature function of liver organoids are still awaited.
As organoids are generated from patient-specific cell sources, liver organoids from patients with DILI may be an invaluable resource for idiosyncratic DILI prediction. Large-scale organoid banks, including healthy donors with their genetic information, can help increase the prediction of human responses by providing adequate statistical power. Additionally, the integration of single-cell RNA sequencing technology and artificial intelligence computational prediction can aid the discovery of the underlying mechanism of complex DILI and biomarkers of drug toxicity. Taken together, advanced human liver models may help to minimize DILI and, their application may be further advanced by pharmaceutical companies and for regulatory purposes.
Notes
Conflict of interest
No potential conflict of interest relevant to this article was reported.
Funding
This work was supported by the Korea Research Institute of Bioscience and Biotechnology (KRIBB) Research Initiative Program (KGM4722223); by a grant (22213MFDS386) from the Ministry of Food and Drug Safety, Korea, in 2022; the National Research Foundation (NRF) grant funded by the Korean government (MSIT) (NRF-2022R1A2B5B02001644).
Author contributions
Conceptualization: SJM, JL, YS, MJS; Writing-original draft preparation: SJM, MJS; Writing-review and editing: SJM, JL, YS, MJS.
Data availability
Please contact the corresponding author for data availability.

